How to plot MODFLOW Head Output in Paraview with Python - Tutorial
/MODFLOW computes the groundwater heads over a porous / fractured media upon a series of boundary conditions as recharge, evapotranspiration, drains, well and others on steady and transient conditions. Free and commercial software is available for the MODFLOW model construction and MODFLOW output representation. Despite the fact the capabilities of these softwares, there are some gaps in data processing and representation; isometric views, animation and custom cross sections are still difficult to achieve under the existing tools, specially on multilayered models with transient conditions over series of time steps and stress periods.
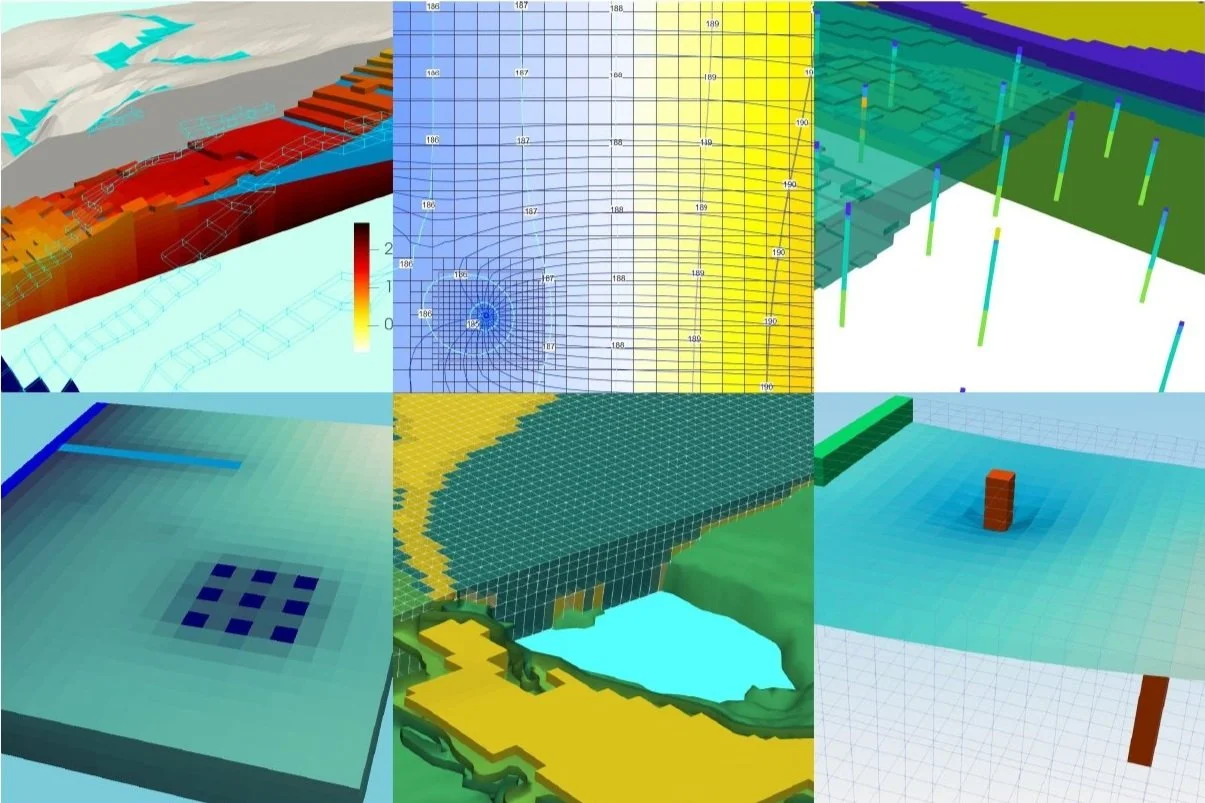
There is a particular open source software for data representation that is of our interest, it is called Paraview (paraview.org). This visual application was designed to analyze extremely large datasets using distributed memory computing resources, in fact the term "para" in Paraview comes from the parallelization of computer cores.
This tutorial shows the complete procedure to create a Paraview compatible geometry data format called *.vtk, and the representation on Paraview. The scripting was done in Python 3 on a Jupyter Notebook. Model input files, output files and project files in Model Muse are available at the end of this article.
Tutorial
Code
This is the complete Python code used in this tutorial:
#import the required libraries
import os, re
import numpy as np
#change directory to the model files path
os.chdir('C:/Users\Saul\Documents\Ih_PlayaroundwithVTK\Model')
#open the *.DIS file and get the model boundaries and discretization
dis= open('Modelo1.dis').readlines()
lowerleft, upperright = re.sub('\(|\)' , ',' , dis[2]), re.sub('\(|\)' , ',' , dis[3])
lowerleft, upperright = lowerleft.split(','), upperright.split(',')
#border coordinates
xmin, ymin = float(lowerleft[1]), float(lowerleft[2])
xmax, ymax = float(upperright[1]), float(upperright[2])
#number of layers, rows and columns
nlay, nrows, ncols = int(dis[6].split()[0]), int(dis[6].split()[1]), int(dis[6].split()[2])
#grid resolution
xres, yres = (xmax-xmin)/ncols, (ymax-ymin)/nrows
#arreglo de easting y northing
eastings = np.linspace(xmin, xmax, ncols+1, endpoint=True, dtype='float32')
northings = np.linspace(ymin, ymax, nrows+1, endpoint=True, dtype='float32')
#bottom elevation of model
linebottom=[x for x in range(len(dis)) if dis[x][:8]=='CONSTANT']
bottom= float(dis[linebottom[1]].split()[1])
#lines that separate heads from a layer
breakers=[x for x in range(len(dis)) if dis[x][:8]=='INTERNAL']
#intervals among breakers
intervals = []
for i in range(len(breakers))[1:]:
delta=breakers[i]-breakers[i-1]
intervals.append(delta)
interval = max(set(intervals))
#counter plus a empty numpy array for model results
i=0
zmatrix = np.zeros((nlay+1,nrows+1,ncols+1))
#loop to read all the heads in one layer, reshape them to the grid dimensions and add to the numpy array
for salto in breakers[1:]:
rowinicio = salto + 1
rowfinal = salto + interval #here we insert the len of lines per layer
matrixlist = []
for linea in range(rowinicio,rowfinal,1):
listaitem = [float(item) for item in dis[linea].split()]
for item in listaitem: matrixlist.append(item)
matrixarray = np.asarray(matrixlist)
matrixarray = matrixarray.reshape(nrows,ncols)
zmatrix[i,:-1,:-1]=matrixarray
i+=1
#add the bottom to the zmatrix
zmatrix[nlay,:,:]=np.ones((nrows+1,ncols+1))*bottom
#complete the extreme right column and lowest row as a duplicate of the previous one
zmatrix[:,-1,:]=zmatrix[:,-2,:]
zmatrix[:,:,-1]=zmatrix[:,:,-2]
#definition of xyz points for all vertex
VTU_Points = []
for lay in range(nlay+1):
for row in range(nrows+1):
for col in range(ncols+1):
xyz=[eastings[col],northings[row],zmatrix[lay, row, col]]
#print(eastings[col],northings[row],zmatrix[lay, row, col])
VTU_Points.append(xyz)
#empty list to store cell coordinates
listahexahedrons = []
maximos = []
#get the nodes and rows per layer
nodesxlay, nodesxrow = (ncols+1)*(nrows+1),ncols+1
#definition of cell coordinates
for lay in range(nlay):
for row in range(nrows):
for col in range(ncols):
pt0 = nodesxlay*(lay+1)+nodesxrow*(row+1)+col
pt1 = nodesxlay*(lay+1)+nodesxrow*(row+1)+col+1
pt2 = nodesxlay*(lay+1)+nodesxrow*(row)+col+1
pt3 = nodesxlay*(lay+1)+nodesxrow*(row)+col
pt4 = nodesxlay*(lay)+nodesxrow*(row+1)+col
pt5 = nodesxlay*(lay)+nodesxrow*(row+1)+col+1
pt6 = nodesxlay*(lay)+nodesxrow*(row)+col+1
pt7 = nodesxlay*(lay)+nodesxrow*(row)+col
lista = [pt0,pt1,pt2,pt3,pt4,pt5,pt6,pt7]
listahexahedrons.append(lista)
#open the *.FHD file to read the heads, beware that the script runs only on static models
hds = open('Modelo1.fhd').readlines()
#lines that separate heads from a layer
breakers=[x for x in range(len(hds)) if hds[x][:4]==' ']
#counter plus a empty numpy array for model results
i=0
listaheads = []
#loop to read all the heads in one layer, reshape them to the grid dimensions and add to the numpy array
for salto in breakers:
rowinicio = salto + 1
rowfinal = salto + 2545 #here we insert the len of lines per layer
matrixlist = []
for linea in range(rowinicio,rowfinal,1):
listaitem = [float(item) for item in hds[linea].split()]
for item in listaitem: listaheads.append(item)
i+=1
#filter hexahedrons and heads for active cells
listahexahedronsdef = []
listaheadsdef = []
for i in range(len(listaheads)):
if listaheads[i] > -1e-10:
listahexahedronsdef.append(listahexahedrons[i])
listaheadsdef.append(listaheads[i])
import os
os.chdir('C:/Users\Saul\Documents\Ih_PlayaroundwithVTK')
from pyvtk import *
points = VTU_Points
vtk = VtkData( UnstructuredGrid(points,
hexahedron=listahexahedronsdef,
),
CellData(Scalars(listaheadsdef)),
'Unstructured Grid Example'
)
vtk.tofile('VTUFiles/VTU_Test2b')
Input files
You can download the input files here.